南京科佰生物科技有限公司官方网站欢迎您的光临!
免费咨询热线: 4008-750-250
科研细胞
药靶细胞
标准品
NGS靶向捕获探针
KIF5B(E15)-RET(E12)-Short/BaF3
| CBP73205 | |
| I. Introduction | |
| Cell Line Name: | KIF5B(E15)-RET(E12)-Short/BaF3 |
| Host Cell: | Ba/F3 |
| Stability: | 16 passages (in-house test, that not means the cell line will be instable beyond the passages we tested.) |
| Application: | Anti-proliferation assay and PD assay |
| Freeze Medium: | 90% FBS+10% DMSO |
| Complete Culture Medium: | RPMI-1640+10%FBS+1 ug/ml puromycin |
| Mycoplasma Status: | Negative |
| II.Background | |
|
Chromosomal rearrangements involving the gene that encodes the RET tyrosine kinase are known oncogenic drivers in 1% to 2% of patients with non–small cell lung cancer (NSCLC). These RET rearrangements occur with characteristic partners, most commonly KIF5B, but also CCDC6, NCOA, TRIM33, CUX1, KIAA1217, FRMD4A, and KIAA1468. They are typically identified in young patients with adenocarcinoma histology and minimal smoking history. Therapeutic targeting of RET-fusion–driven NSCLCs has taken the form of treatment with broad-spectrum tyrosine kinase inhibitors with anti-RET activity, such as cabozantinib (Cabometyx; Cometriq), vandetanib (Caprelsa), lenvatinib (Lenvima), RXDX-105, and sunitinib (Sutent). Cabozantinib and vandetanib have been the most heavily studied multi-kinase inhibitors (MKIs), with response rates of 20% to 50% in largely pretreated patients with RET-rearranged NSCLC. Sunitinib has been used in fewer patients to date with initial results demonstrating a 22% response rate. RXDX-105 has exhibited uniquely impressive response rates (75%) in patients with non–KIF5B-RET-fusion NSCLC, compared with 0% response in patients with KIF5B-RET-fusion–positive NSCLC. BLU-667 has demonstrated an objective response rate of 50% in patients with RET-fusion positive NSCLC, and LOXO-292 reported a 74% ORR in patients with RET-fusion positive NSCLC. Notably, RXDX-105, BLU- 667, and LOXO-292 have all demonstrated some central nervous system activity in these early phase trials. Future directions of RET inhibition in patients with RET-rearranged NSCLC include additional clinical validation of the next generation RET-selective inhibitors RXDX-105, BLU-667, and LOXO-292 and comparing multikinase inhibitors with RET-selective inhibitors to determine the optimal sequencing of RET-targeted therapies. |
|
| III. Representative Data | |
|
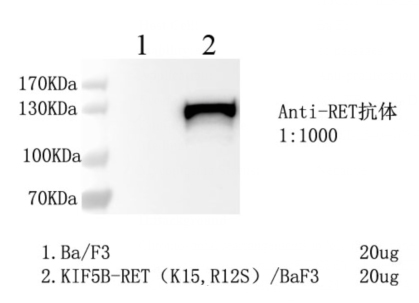
1. WB of KIF5B-RET (K15, R12S)/BaF3
|
|
|
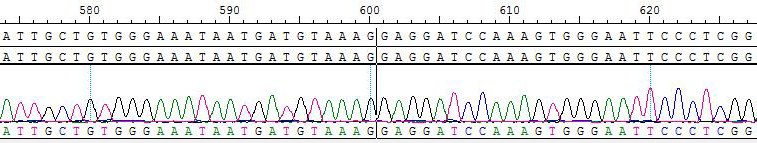
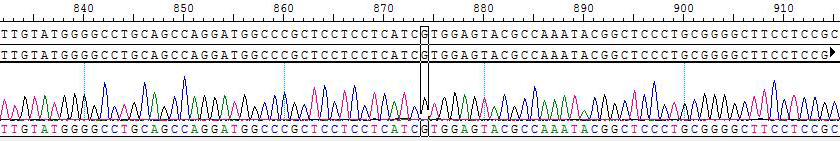
2. Sanger of KIF5B-RET (K15, R12S)/BaF3
|
|
|
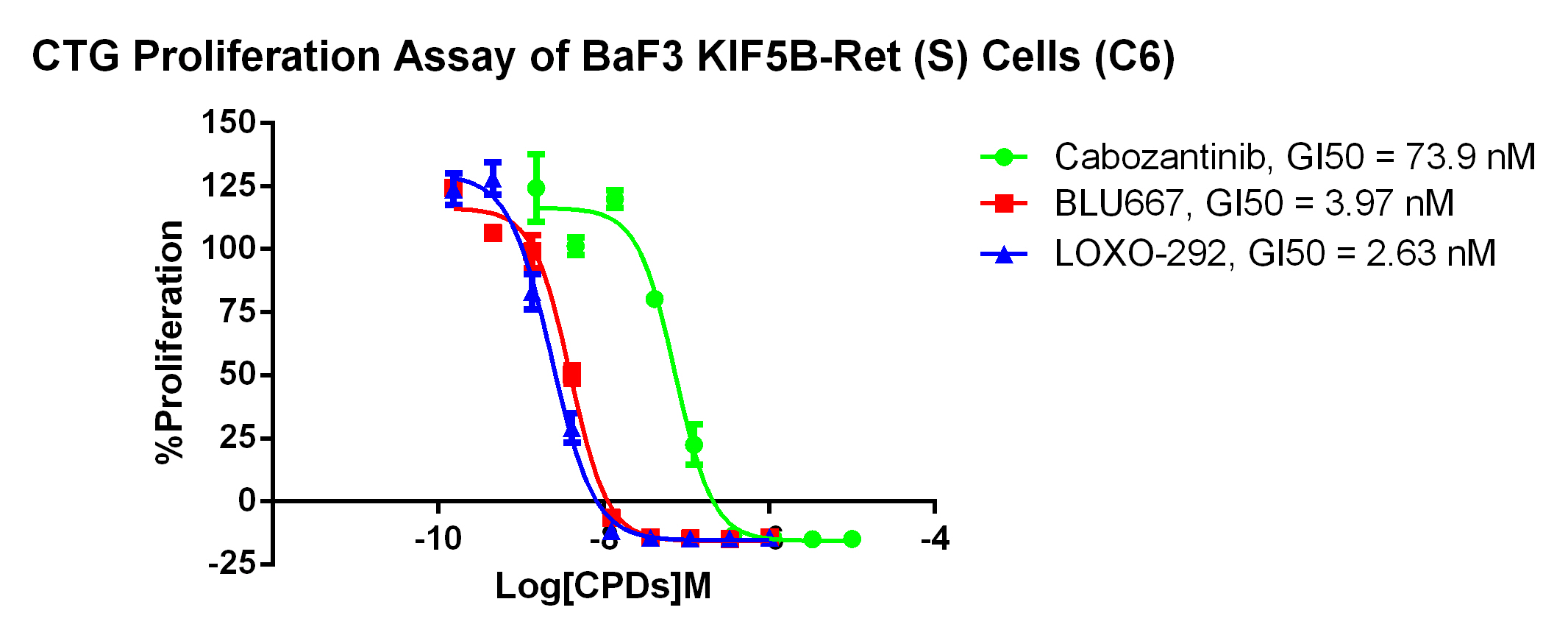
3. Anti-proliferation assay
Figure 4. CTG Proliferation Assay of BaF3 KIF5B-Ret (s) Cells (C6). |
|

扫一扫,关注我们公众号
Copyright ©2021 www.cobioer.com All Rights Reserved. 南京科佰生物科技有限公司版权所有 苏ICP备13038923号